Note
Go to the end to download the full example code or to run this example in your browser via Binder
Scikit-fda and scikit-learn#
In this section, we will explain how scikit-fda interacts with the popular machine learning package scikit-learn. We will introduce briefly the main concepts of scikit-learn and how scikit-fda reuses the same concepts extending them to the functional data analysis field.
# Author: Carlos Ramos Carreño
# License: MIT
A brief summary of scikit-learn architecture#
The library scikit-learn is probably the most
well-known Python package for machine learning. This package focuses in
machine learning using multivariate data, which should be stored in a numpy
ndarray in order to process it. However, this library has
defined a particular architecture that can be followed in order to provide
new tools that work in situations not even imagined by the original authors,
while remaining compatible with the tools already provided in scikit-learn.
In scikit-fda, the same architecture is applied in order to work with
functional data observations. As a result, scikit-fda tools are
largely compatible with scikit-learn tools, and it is possible to reuse
objects such as pipelines or even
hyperparameter selection methods such as
grid search cross-validation
in the functional data setting.
We will introduce briefly the main concepts in scikit-learn, and explain how the tools in scikit-fda are related with them. This is not intended as a full explanation of scikit-learn architecture, and the reader is encouraged to look at the scikit-learn tutorials in order to achieve a deeper understanding of it.
The Estimator object#
A central concept in scikit-learn (and scikit-fda) is what is called an estimator. An estimator in this context is an object that can learn from the data. Thus, classification, regression and clustering methods, as well as transformations with parameters learned from the training data are particular kinds of estimators. Estimators can also be instanced passing parameters, which can be tuned to the data using hyperparameter selection methods.
Estimator objects have a fit method, with receive the training data
and (if necessary) the training targets. This method uses the training data
in order to learn some parameters of a model. When the learned parameters
are part of the user-facing API, then by convention they are attributes of
the estimator ending in with the _ character.
As a concrete example of this, consider a nearest centroid classifier
for functional data. The object
NearestCentroid is a classifier, and
thus an estimator. As part of the training process the centroids of
the classes are computed and available as the learned parameter
centroids_.
Note
The function train_test_split() is
one of the functions originally from scikit-learn that can be
directly reused in scikit-fda.
import skfda
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
X, y = skfda.datasets.fetch_growth(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
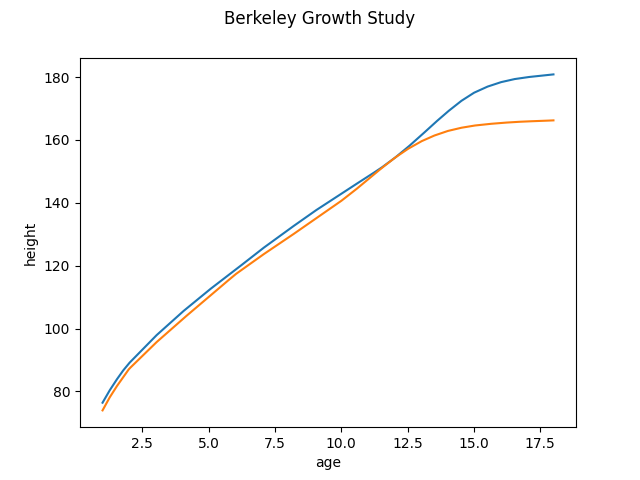
classifier = skfda.ml.classification.NearestCentroid()
classifier.fit(X_train, y_train)
classifier.centroids_.plot()
plt.show()
Transformers#
Transformers are estimators which can convert
data to a new form. Examples of them are preprocessing methods, such as
smoothing, registration and dimensionality reduction methods. They always
implement fit_transform for fitting and transforming the data in one
step. The transformers may be inductive, which means that
can transform new data using the learned parameters. In that case they
implement the transform method to transform new data. If the
transformation is reversible, they usually also implement
ìnverse_transform.
As an example consider the smoothing method
skfda.preprocessing.smoothing.NadarayaWatsonHatMatrix. Smoothing
methods attempt to remove noise from the data leveraging its continuous
nature.
As these methods discard information of the original data they usually are
not reversible.
import skfda.preprocessing.smoothing as ks
from skfda.misc.hat_matrix import NadarayaWatsonHatMatrix
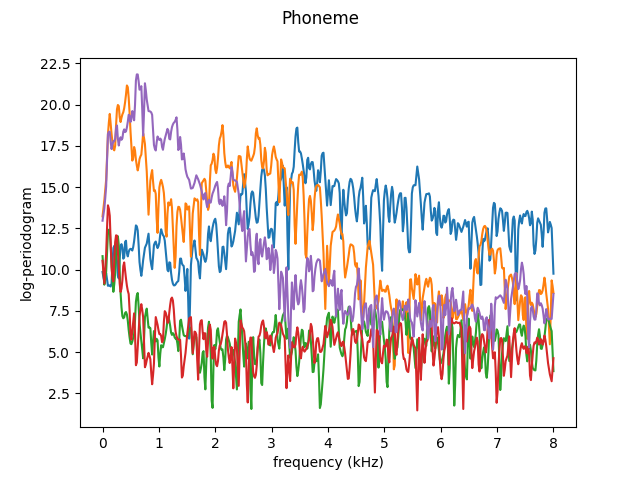
X, y = skfda.datasets.fetch_phoneme(return_X_y=True)
# Keep the first 5 functions
X = X[:5]
X.plot()
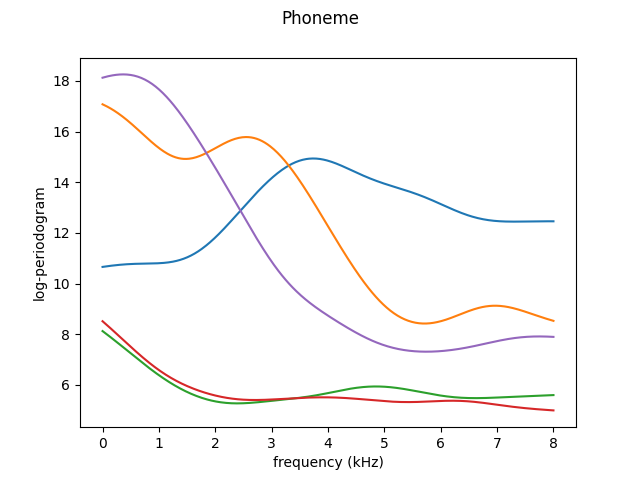
smoother = ks.KernelSmoother(kernel_estimator=NadarayaWatsonHatMatrix())
X_smooth = smoother.fit_transform(X)
X_smooth.plot()
plt.show()
Predictors (classifiers, regressors, clusterers…)#
Predictors in scikit-learn are estimators that can assign a certain target to a particular observation. This includes supervised methods such as classifiers (for which the target will be a class label), or regressors (for which the target is a real value, a vector, or, in functional data analysis, even a function!) and also unsupervised methods such as clusterers or outlying detector methods.
Predictors should implement the fit_predict method for fitting the
estimators and predicting the targets in one step and/or the predict
method for predicting the targets of possibly non previously observed data.
Usually transductive estimators implement only the former
one, while inductive estimators implement the latter one (or
both).
Predictors can have additional non-mandatory methods, such as
predict-proba for obtaining the probability of a particular prediction
or score for evaluating the results of the prediction.
As an example, we can look at the KMeans
clustering method for functional data. This method will try to separate
the data into different clusters according to the distance between
observations.
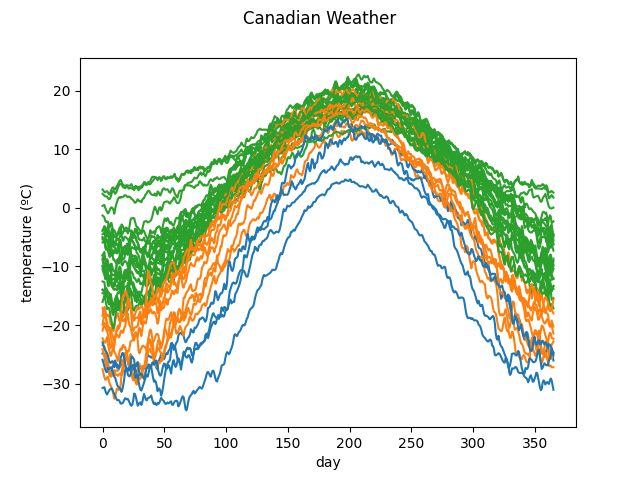
X, y = skfda.datasets.fetch_weather(return_X_y=True)
# Use only the first value (temperature)
X = X.coordinates[0]
clusterer = skfda.ml.clustering.KMeans(n_clusters=3)
y_pred = clusterer.fit_predict(X)
X.plot(group=y_pred)
plt.show()
Metaestimators#
In scikit-learn jargon, a metaestimator is an estimator that takes other estimators as parameters. There are several reasons for doing that, which will be explained now.
Composition metaestimators#
It is very common in machine learning to apply one or more preprocessing
steps one after the other, before applying a final predictor. For this
purpose scikit-learn offers the Pipeline, which
join the steps together and uses the same estimator API for performing all
steps in order (this is usually referred as the composite pattern in
software engineering). The Pipeline estimator
can be used with the functional data estimators available in scikit-fda.
Moreover, as transformers such as dimensionality reduction methods can
convert functional data to multivariate data usable by scikit-learn methods
it is possible to mix methods from scikit-fda and scikit-learn in the same
pipeline.
Warning
In addition, scikit-learn offers estimators that can join several
transformations as new features of the same dataset (
FeatureUnion) or that can apply different
transformers to different columns of the data
(ColumnTransformer). These transformers
are not yet usable with functional data.
As an example, we can construct a pipeline that registers the data using shift registation, then applies a variable selection method to transform each observation to a 3D vector and then uses a SVM classifier to classify the data.
from skfda.preprocessing.dim_reduction import variable_selection as vs
from skfda.preprocessing.registration import LeastSquaresShiftRegistration
from sklearn.pipeline import Pipeline
from sklearn.svm import SVC
X, y = skfda.datasets.fetch_growth(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
pipeline = Pipeline([
("registration", LeastSquaresShiftRegistration()),
("dim_reduction", vs.RKHSVariableSelection(n_features_to_select=3)),
("classifier", SVC()),
])
pipeline.fit(X_train, y_train)
pipeline.score(X_test, y_test)
1.0
Hyperparameter optimizers#
Some of the parameters used for the creation of an estimator need to be
tuned to each particular dataset in order to improve the prediction accuracy
and generalization. There are several techniques to do that already
available in scikit-learn, such as grid search cross-validation
(GridSearchCV) or randomized search
(RandomizedSearchCV). As these
hyperparameter optimizers only need to split the data and call score in
the predictor, they can be directly used with the methods in scikit-fda.
Note
In addition one could use any optimizer that understand the scikit-learn API such as those in scikit-optimize.
As an example, we will use GridSearchCV
to select the number of neighbors used in a
KNeighborsClassifier.
from sklearn.model_selection import GridSearchCV
X, y = skfda.datasets.fetch_growth(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
classifier = skfda.ml.classification.KNeighborsClassifier()
grid_search = GridSearchCV(
estimator=classifier,
param_grid={"n_neighbors": range(1, 10, 2)},
)
grid_search.fit(X_train, y_train)
n_neighbors = grid_search.best_estimator_.n_neighbors
score = grid_search.score(X_test, y_test)
print(n_neighbors, score)
3 0.9583333333333334
Ensemble methods#
The ensemble methods VotingClassifier and
VotingRegressor in scikit-learn use several
different estimators in order to predict the targets. As this is done
by evaluating the passed estimators as black boxes, these predictors can
also be combined with scikit-fda predictors.
Warning
Other ensemble methods, such as
BaggingClassifier or
AdaBoostClassifier cannot yet
be used with functional data unless it has been
transformed to a multivariate dataset.
As an example we will use a voting classifier to classify data using as classifiers a knn-classifier, a nearest centroid classifier and a maximum depth classifier.
from sklearn.ensemble import VotingClassifier
X, y = skfda.datasets.fetch_growth(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
knn = skfda.ml.classification.KNeighborsClassifier()
nearest_centroid = skfda.ml.classification.NearestCentroid()
mdc = skfda.ml.classification.MaximumDepthClassifier()
voting = VotingClassifier([
("knn", knn),
("nearest_centroid", nearest_centroid),
("mdc", mdc),
])
voting.fit(X_train, y_train)
voting.score(X_test, y_test)
0.75
Multiclass and multioutput classification utilities#
The scikit-learn library also offers additional utilities that can convert
a binary classifier into a multiclass classifier (such as
OneVsRestClassifier) or to extend a single
output classifier or regressor to accept also multioutput (vector-valued)
targets.
In this example we want to use as a classifier the combination of a
dimensionality reduction method (
RKHSVariableSelection)
and a SVM classifier (SVC). As that particular
dimensionality reduction method is only suitable for binary data, we use
OneVsRestClassifier to classify in a
multiclass dataset.
from sklearn.multiclass import OneVsRestClassifier
X, y = skfda.datasets.fetch_phoneme(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
pipeline = Pipeline([
("dim_reduction", vs.RKHSVariableSelection(n_features_to_select=3)),
("classifier", SVC()),
])
multiclass = OneVsRestClassifier(pipeline)
multiclass.fit(X_train, y_train)
multiclass.score(X_test, y_test)
0.9140070921985816
Other scikit-learn utilities#
In addition to the aforementioned objects, there are plenty of objects in
scikit-learn that can be applied directly to functional data. We have
already seen in the examples the function
train_test_split(). Other objects and
functions such as KFold can be directly
applied to functional data in order to split it into folds. Scorers for
classification or regression, such as
accuracy_score() can be directly applied to
functional data problems.
Moreover, there are plenty of libraries that aim to extend scikit-learn in several directions (take a look at the list of related projects). You will probably see that a lot of the functionality can be applied to scikit-fda, as it uses the same API as scikit-learn.
Total running time of the script: (0 minutes 6.341 seconds)